This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
Conclusions and Future Work
To give a little background on the traits associated with rs12722, I would first like to define running economy as which can be quantified
by measuring energy utilization while moving at an an aerobic intensity. The variant causes an increase in running economy due to it also causing a decrease in flexibility(1). Inflexibility has been shown to benefit long distance runners. Knowing this, the purpose of this project was to determine how inflexibility was caused.
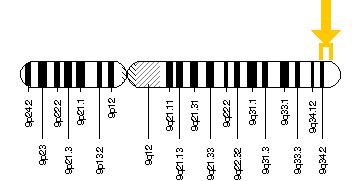
Before I could answer this question, I needed to look into the background to learn more how this variant could result in such a drastic phenotype. This gene is fairly large, with COL5A1 being found on the long arm of chromosome 9 being composed of approximately 200,000 base pairs (2). COL5A1 codes for a protein called pro-alpha 1(V) chain. This forms type V collagen with 3 other sub-units.
Before I could answer this question, I needed to look into the background to learn more how this variant could result in such a drastic phenotype. This gene is fairly large, with COL5A1 being found on the long arm of chromosome 9 being composed of approximately 200,000 base pairs (2). COL5A1 codes for a protein called pro-alpha 1(V) chain. This forms type V collagen with 3 other sub-units.
The mutation is actually the result of a single nucleotide polymorphism (SNP) in the 3' UTR of the COL5A1 gene. The difference between rs12722 and COL5A1 is that instead of the normal cytosine one would find, there is actually a thymine.
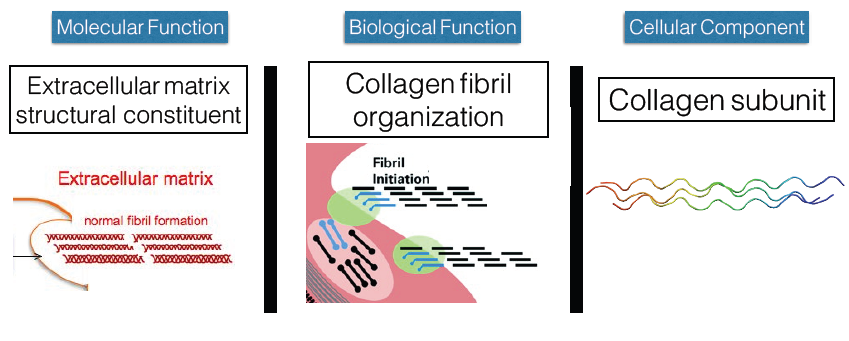
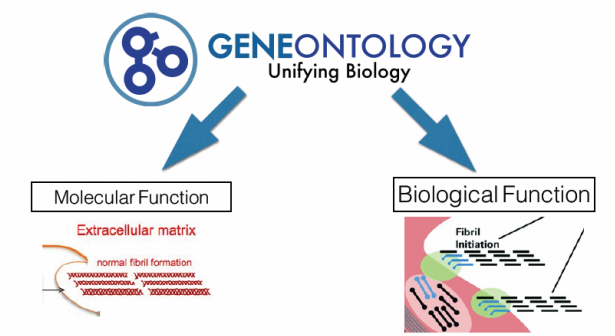
Next I looked at Gene Ontology to determine some of the functions the gene and the protein, pro-alpha 1(V) chain, has that could possibly be related to inflexibility. For the molecular function of the gene, I found it was an extra-cellular constituent, meaning it aided in structural support of different tissues within the cell. For the biological function, I found that it was involved in collagen fibril organization. Further research on this point showed that it was needed to initiate fibril organization and synthesis (4). And finally for the cellular component, it was shown to be a collagen subunit. This of course was to be expected as it is known that three units are needed to form type V collagen as mentioned before.
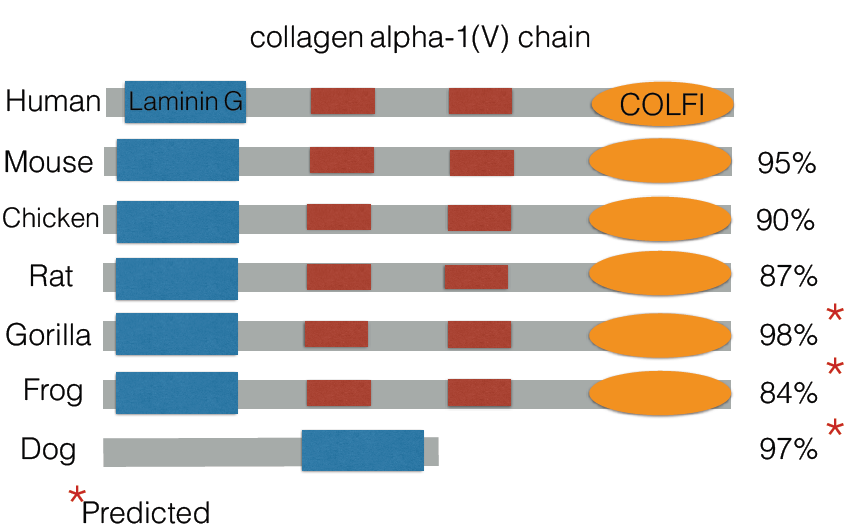
I next looked into the structure of the pro-alpha 1(V) chain to determine if there were any domains that explained the function. I found that it contained three main domains. It contained multiple collagen triple helix repeats, which is a staple of collagen genes. It also contained the COLFI domain. This was found at the C terminus of collagen molecule, but the exact function of this domain is still unknown. It contained a laminin domain as well, which binds to laminin and is important to extracellular structure and support. This makes sense given its function as a structural protein.
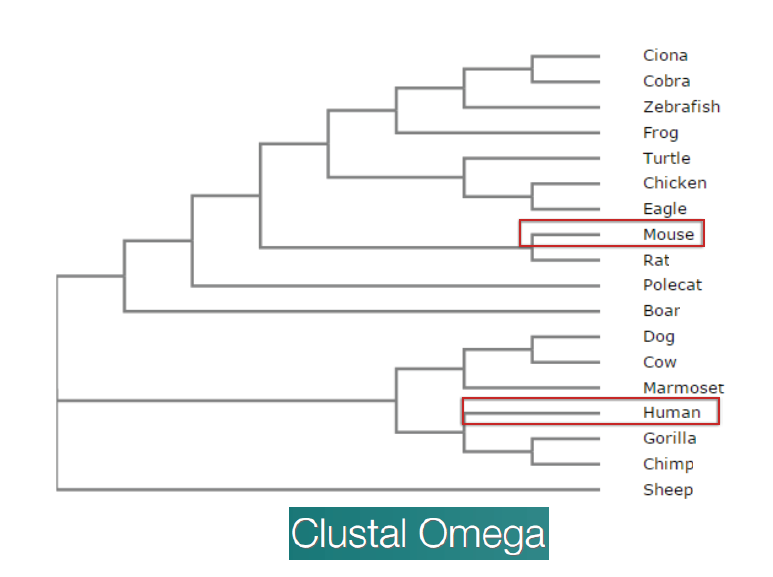
I next looked to see how conserved it was amongst other organisms. From the online databases, there were a lot of organisms that did not have the exact sequence confirmed. Given all the results though, as well as the predictions, the gene is highly conserved in animals. I followed up on this by creating a phylogeny tree of the protein to see conservation amongst different species using Clustal Omega.
Given how similar the proteins were between mice and humans, and how easy mice are to handle, I knew they would be my model organism when pursuing my specific aims.
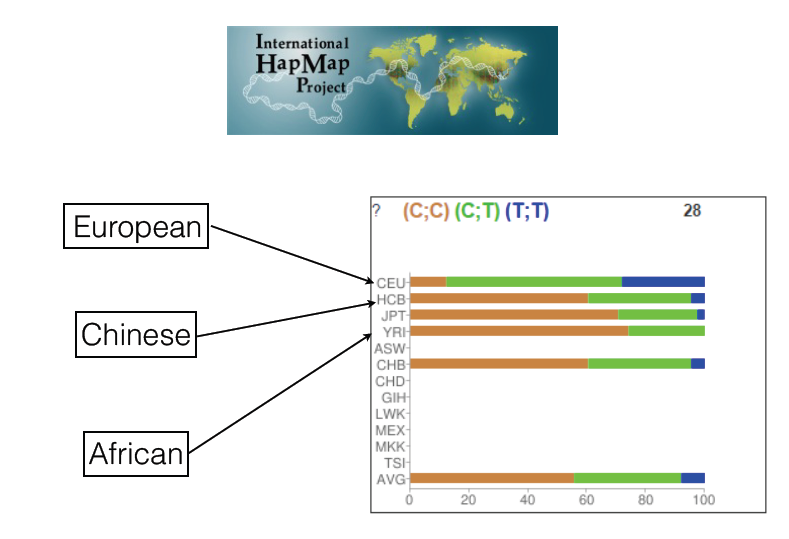
Before looking into those, I was curious to see the distribution of the gene amongst different ethnic backgrounds. Using HapMap, which is essentially a catalog of common genetic mutations throughout different populations, I found one of rs12722. The blue represents the population that is homozygous for rs12722, the green represents those that are heterozygous for rs12722 and COL5A1, and the brown represents those that are homozygous for COL5A1.
Before looking into those, I was curious to see the distribution of the gene amongst different ethnic backgrounds. Using HapMap, which is essentially a catalog of common genetic mutations throughout different populations, I found one of rs12722. The blue represents the population that is homozygous for rs12722, the green represents those that are heterozygous for rs12722 and COL5A1, and the brown represents those that are homozygous for COL5A1.
Looking at the distribution for those who are homozygous for rs12722, 25% of people of European background, 5% of people of Chinese descent, and 0% of those from a tribe in Zimbabwe contain this genotype. It is important to not that this project is still being worked on, so the sample sizes will grow over time.
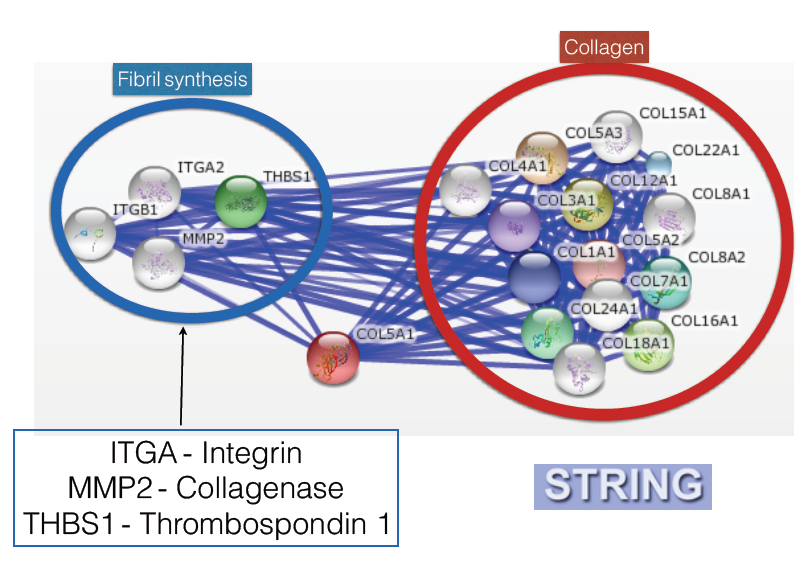
Next I looked at the protein-protein interaction network of the COL5A1 gene product to try to further determine other possible functions that could cause inflexibility. These results were obtained from the database String.
Next I looked at the protein-protein interaction network of the COL5A1 gene product to try to further determine other possible functions that could cause inflexibility. These results were obtained from the database String.
A lot of the shown interactions were to be expected. Pro-alpha 1(V) chain reacts with a lot of different collagens. This makes sense as multiple types of collagen are used to make collagen fibers. There were other interactions that further explained COL5A1's role in fibril synthesis. ITGA codes for Integrin, a protein found on the outside of cells that contains a domain that is used to bind to collagen. Next there was MMP2, which codes for a collagenase. This is used to dissolve collagen fibers. This makes sense as there should be something that can remove these fibrils if necessary. Finally there was THBS, which codes for thrombospondin-1. This is involved in angiogenesis. This also makes sense as blood vessels are made of connective tissues, something that is composed of collagen.
So given this information, it is still unknown the rs12722 gene product differs from the normal
COL5A1 gene product so that cellular functionality is affected and an enhanced running economy is produced. I hypothesize that the rs12722 gene product enhances running economy due to an increase in fibril formation as a result of higher production of type V collagen as a result of more pro-alpha 1(V) chain. This will promote extracellular matrix support and therefore create more muscle inflexibility. This in turn enhances running economy. By testing my specific aims, I hope to confirm this hypothesis.
So given this information, it is still unknown the rs12722 gene product differs from the normal
COL5A1 gene product so that cellular functionality is affected and an enhanced running economy is produced. I hypothesize that the rs12722 gene product enhances running economy due to an increase in fibril formation as a result of higher production of type V collagen as a result of more pro-alpha 1(V) chain. This will promote extracellular matrix support and therefore create more muscle inflexibility. This in turn enhances running economy. By testing my specific aims, I hope to confirm this hypothesis.
For my first specific aim, I wish to determine if the SNP is within a DNA motif or if a miRNA
exists that would bind with the COL5A1 mRNA at an area that would contain the SNP if it was a rs12722 mRNA transcript. I predict that the mutation is present on a regulatory DNA binding
domain or affects the binding of a miRNA to the normal mRNA transcript, which would regulate translation. I believe this could be the case because miRNA's tend to bind to the 3' UTR region of mRNA transcripts to regulate translation.
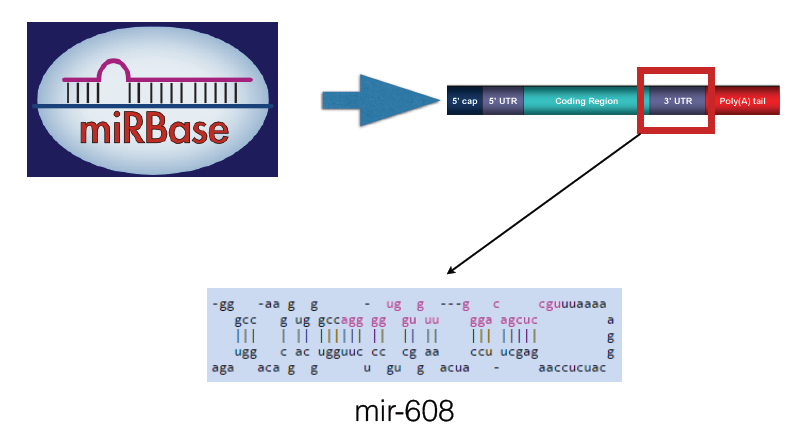
I used DREME to test for DNA motifs and miRBase to look for possible miRNA sequences that could bind to the COL5A1 mRNA transcript. While I did not find any motifs for the DNA, I found a some miRNAs that bind pretty close to the area where the SNP would be in rs12722. I later found a paper that found that another miRNA binds to the normal COL5A1 transcript and that there is strong evidence that this mutation alters mRNA structure to reduce miRNA regulation (5). The miRNA is characterized as mir-608.
exists that would bind with the COL5A1 mRNA at an area that would contain the SNP if it was a rs12722 mRNA transcript. I predict that the mutation is present on a regulatory DNA binding
domain or affects the binding of a miRNA to the normal mRNA transcript, which would regulate translation. I believe this could be the case because miRNA's tend to bind to the 3' UTR region of mRNA transcripts to regulate translation.
I used DREME to test for DNA motifs and miRBase to look for possible miRNA sequences that could bind to the COL5A1 mRNA transcript. While I did not find any motifs for the DNA, I found a some miRNAs that bind pretty close to the area where the SNP would be in rs12722. I later found a paper that found that another miRNA binds to the normal COL5A1 transcript and that there is strong evidence that this mutation alters mRNA structure to reduce miRNA regulation (5). The miRNA is characterized as mir-608.
To confirm this more sequencing of miRNA and more deep sequencing of mRNA must be done to confirm this interaction.
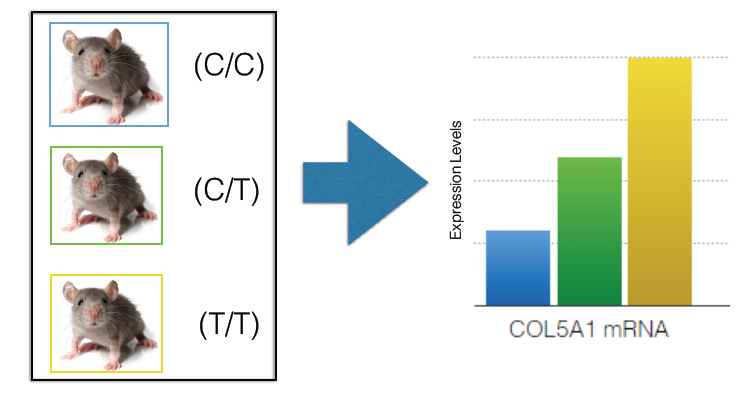
For the second aim, I wanted to determine if there are higher levels of rs12722 mRNA compared to normal COL5A1 mRNA in smooth muscle cells. I wanted to look at smooth muscle cells since previous genetic expression data showed that that is where it is highly expressed. I believe that more rs12722 mRNA will be quantified, which implies creation of more collagen fibrils. To do this, I would create different lines of mice containing the normal human COL5A1 and the rs12722 variant and use RNA sequencing to quantify mRNA levels. I would create 3 lines of mice: homozygous mutant mice, heterozygous mice, and homozygous wild type mice. Then using RNA Sequencing of smooth muscle tissues of the mice.
For the second aim, I wanted to determine if there are higher levels of rs12722 mRNA compared to normal COL5A1 mRNA in smooth muscle cells. I wanted to look at smooth muscle cells since previous genetic expression data showed that that is where it is highly expressed. I believe that more rs12722 mRNA will be quantified, which implies creation of more collagen fibrils. To do this, I would create different lines of mice containing the normal human COL5A1 and the rs12722 variant and use RNA sequencing to quantify mRNA levels. I would create 3 lines of mice: homozygous mutant mice, heterozygous mice, and homozygous wild type mice. Then using RNA Sequencing of smooth muscle tissues of the mice.
I would expect more rs12722 mRNA transcripts to be present. If that were the case, I would follow up with a mouse treadmill experiment to see if those that had were homozygous for rs12722 could run for a longer distance. So overall I would expect that there would be higher levels of of rs12722 mRNA compared to normal COL5A1 mRNA and mice homozygous for rs12722 can for run longer.
For the third aim, I would want to determine if there are any new proteins that interact with the rs12722 variant gene product. I expect that there is a new protein interaction that promotes collagen fibril initiation or is an extracellular matrix structural constituent. By using a TAP tag on the rs12722 variant, I can see if there are any noticeable or new protein interactions with the rs12722 protein that are not present in normal pro-alpha-1(V) chain. I can compare the results to the current known protein to protein interaction network of pro-alpha-1(V) chain as a control.If there were new interactions, I would use Gene Ontology to see the functions of the proteins.
For the third aim, I would want to determine if there are any new proteins that interact with the rs12722 variant gene product. I expect that there is a new protein interaction that promotes collagen fibril initiation or is an extracellular matrix structural constituent. By using a TAP tag on the rs12722 variant, I can see if there are any noticeable or new protein interactions with the rs12722 protein that are not present in normal pro-alpha-1(V) chain. I can compare the results to the current known protein to protein interaction network of pro-alpha-1(V) chain as a control.If there were new interactions, I would use Gene Ontology to see the functions of the proteins.
I would expect these new proteins promote collagen fibril organization or would be an extracellular matrix structural constituent.
For future directions to take, larger studies from marathon participants could be performed to further provide evidence for the correlation between rs12722, inflexibility, and enhanced running economy. Next, more sequencing of miRNAs can be done to confirm the existence of possible translation regulators of normal COL5A1 mRNA transcripts. Another follow up could be A RT-qPCR experiment on mRNA transcription levels in the different line of mice mentioned earlier to confirm the exact expression levels. Finally, a yeast two-hybrid specific library approach could be used to further look for new protein-protein interactions that might not exist for the normal COL5A1 gene product.
For future directions to take, larger studies from marathon participants could be performed to further provide evidence for the correlation between rs12722, inflexibility, and enhanced running economy. Next, more sequencing of miRNAs can be done to confirm the existence of possible translation regulators of normal COL5A1 mRNA transcripts. Another follow up could be A RT-qPCR experiment on mRNA transcription levels in the different line of mice mentioned earlier to confirm the exact expression levels. Finally, a yeast two-hybrid specific library approach could be used to further look for new protein-protein interactions that might not exist for the normal COL5A1 gene product.
References
- COLLAGEN GENE SEQUENCE VARIANTS IN EXERCISE-RELATED TRAITS. Central European Journal of Sport Sciences and Medicine | Vol. 1, No. 1/2013: 3–17
- (2006) COL5A1 gene. Genetics Home Reference. http://ghr.nlm.nih.gov/gene/COL5A1
- http://www.snpedia.com/index.php/Rs12722
- Type V collagen: molecular structure and fibrillar organization of the chicken alpha 1(V) NH2-terminal domain, a putative regulator of corneal fibrillogenesis.The Journal of Cell Biology. 1993;121(5):1181-1189.
- Polymorphisms within the COL5A1 3'-UTR that alters mRNA structure and the MIR608 gene are associated with Achilles tendinopathy. Ann Hum Genet. 2013 May;77(3):204-14. doi: 10.1111/ahg.12013. Epub 2013 Jan 24.
| Final COL5A1 presentation.pdf | |
| File Size: | 3462 kb |
| File Type: | |
| col5a1_presentation.pdf | |
| File Size: | 1673 kb |
| File Type: | |
| logansilberpresentationrd2.pdf | |
| File Size: | 2102 kb |
| File Type: | |
Website Author: Logan Silber
Contact email: [email protected]
Page last updated: 5/15/2015
Course Homepage: www.genetics564.weebly.com
Contact email: [email protected]
Page last updated: 5/15/2015
Course Homepage: www.genetics564.weebly.com